Introducing inSēquio™
The First Programmable 3D CAD Tool for DNA Nanostructure Design-
See our
preprint
on

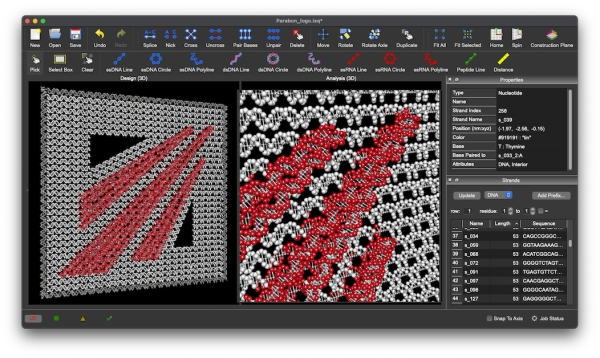
At Parabon, we're pioneering the future of DNA nanotechnology with our advanced inSēquio Design Studio software and comprehensive custom design and synthesis services. Whether you're diving into DNA design with our groundbreaking software or leveraging our expert team for bespoke solutions, we can help you accelerate your nanotechnology research.
Work Across Multiple Dimensions
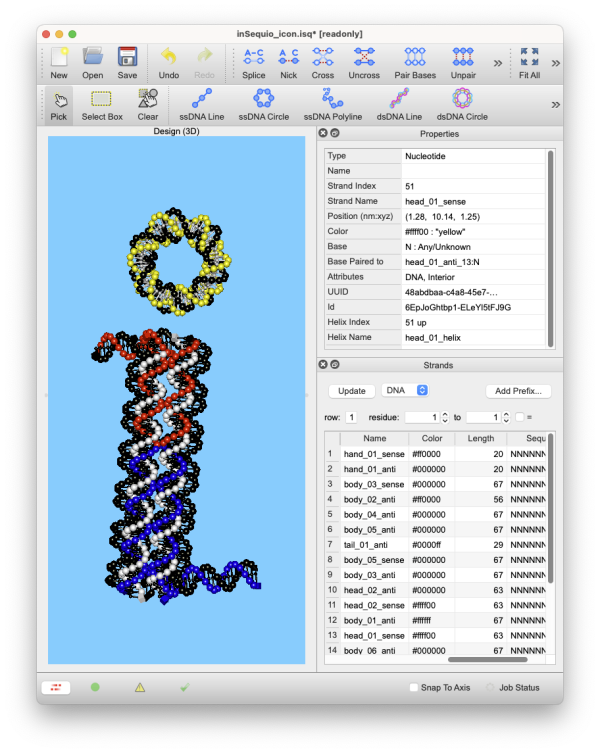
Draft 2D blueprints, sculpt 3D structures, or work with both at the same time!
inSēquio's Schematic (2D), Design (3D), and Analysis (3D) views allow simultaneous editing and highlighting that provide clarity and insight for even the most complex designs.
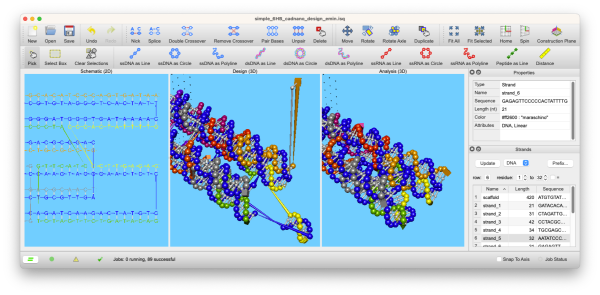
Codify Your Intent
Let's face it. Constructing large, complex designs by hand is time consuming, tedious and error prone. Use inSēquio's Python API to create exactly the design you want in a fast and reproducible way.
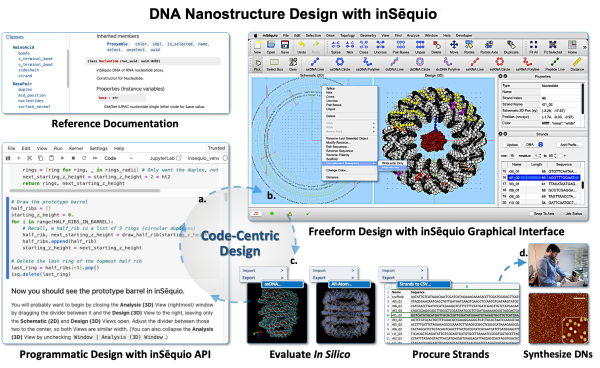
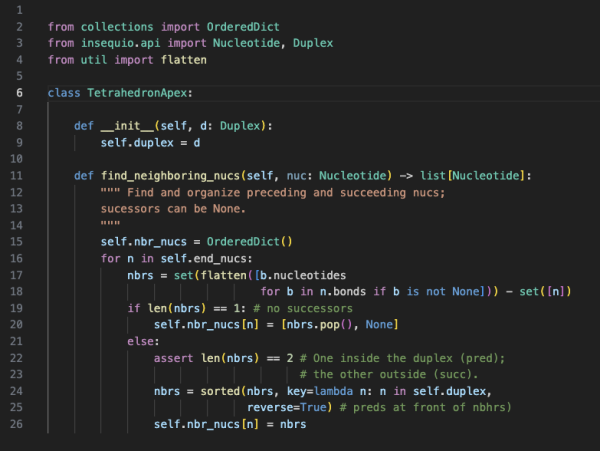
Get Under the Hood
Sometimes you have to get your hands dirty. Zoom in close and use inSēquio's many manipulation tools to tweak, nudge, jigger, rotate, or otherwise force DNA to bend exactly to your will.
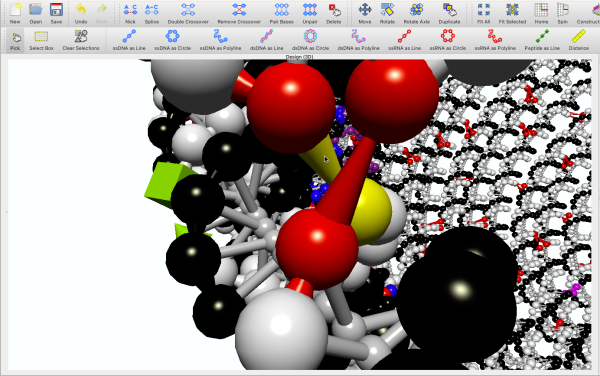
Mashup — Molecularly
DNA is at its most exciting when other elements are involved. Whether you are designing drugs, sensors, or a fleet of nanoscale robots, inSēquio lets you incorporate a wide array of molecular components into your creation.
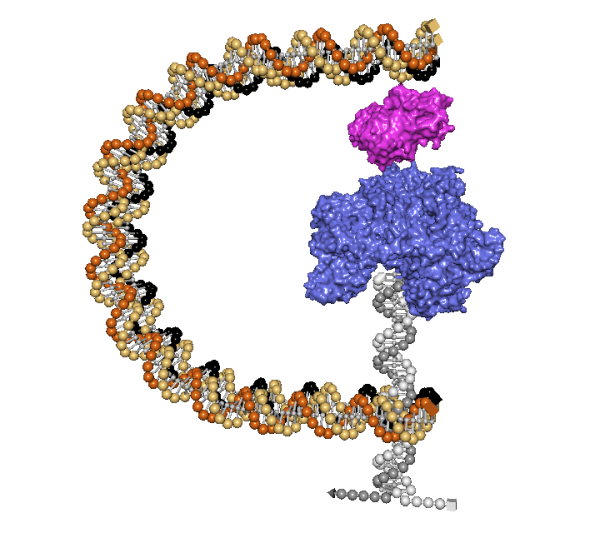
Design Freely
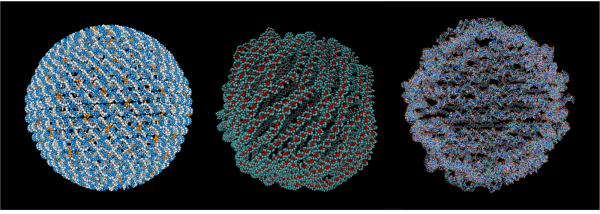
Lattice and wireframe designs comprise the vast majority of all DNA nanostructures ever created, but there is practically unlimited design space that has yet to be explored. Liberate your creativity with inSēquio's free-form design tools and bring forth nanostructures the likes of which the world has never seen.

Simulation Simplified
Launch energy minimization or oxDNA simulations directly from inSēquio and run them in the cloud. Prefer local control? Export all-atom configurations and execute them on your own system. Either way, experience hassle-free, insightful feedback to refine your designs swiftly.
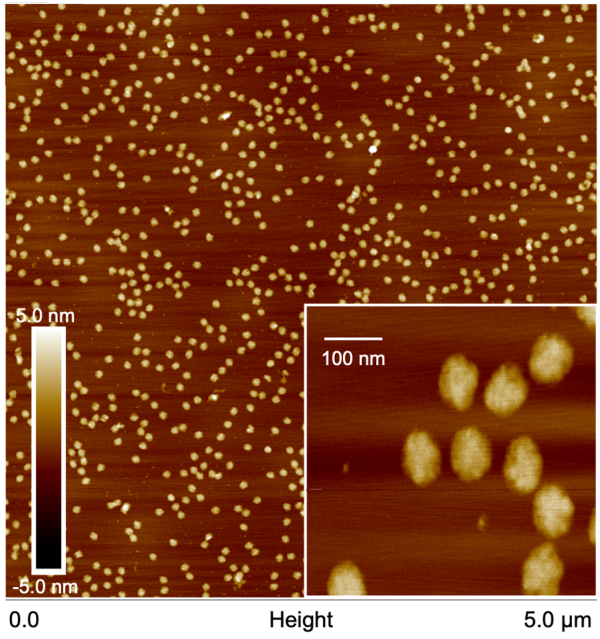
Accelerate Your Research
Whether you are racing to get a product to market or trying to secure more high-impact publications, productivity is the key to reaching your goals before your competition.
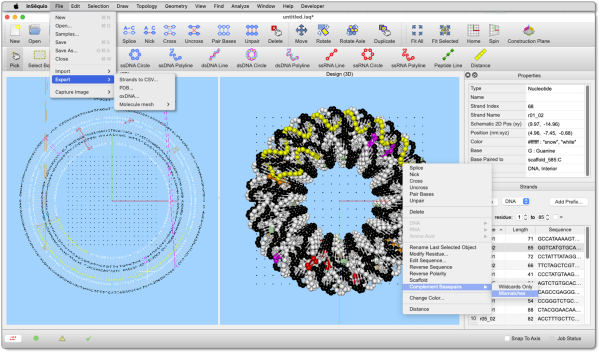
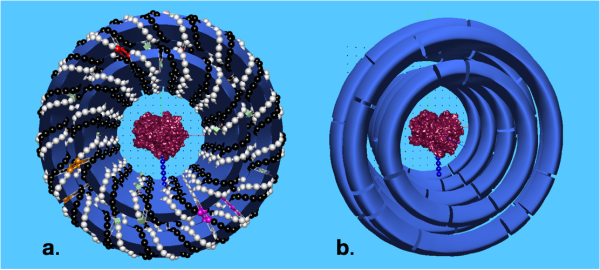
Partner with Us for Groundbreaking Research
Are you looking to make a significant impact in your research field? We're here to help! We have a robust track record in securing research grants and building successful collaborations with both academic and industry leaders. Let’s innovate together and achieve remarkable results.
Contact us today to discuss how we can collaborate on future projects and leverage our mutual strengths for game-changing discoveries.
For more information, please email insequio-sales@parabon.com or call (703) 689-9689 x207


